Today, a friend forwarded me this opinion piece about the murder-suicide of NFL linebacker Jovan Belcher. I responded that I blogged about such cases before, and I didn't intend to write yet another blog post about chronic traumatic encephalopathy. I felt as though lethal violence among football players is happening so often now, it's not even worth yet another post. But, the fact that the violence has reached this level of banality is in itself worthy of comment. Don't worry, though, the next time an NFL player commits homicide or suicide, I promise not to write about it.
I discuss issues pertaining to the practice of neuropathology -- including nervous system tumors, neuroanatomy, neurodegenerative disease, muscle and nerve disorders, ophthalmologic pathology, neuro trivia, neuropathology gossip, job listings and anything else that might be of interest to a blue-collar neuropathologist.
Wednesday, June 26, 2013
Best Post of December 2012: "the NFL is a breeding ground for mental illness" -- Former Denver Bronco Nate Jackson
Today, a friend forwarded me this opinion piece about the murder-suicide of NFL linebacker Jovan Belcher. I responded that I blogged about such cases before, and I didn't intend to write yet another blog post about chronic traumatic encephalopathy. I felt as though lethal violence among football players is happening so often now, it's not even worth yet another post. But, the fact that the violence has reached this level of banality is in itself worthy of comment. Don't worry, though, the next time an NFL player commits homicide or suicide, I promise not to write about it.
Saturday, June 15, 2013
Mutations in COQ2 in Familial and Sporadic Multiple System Atrophy
Researchers from the Multiple System Atrophy (MSA) Research Collaborative in Japan just published online in the New England Journal of Medicine findings providing evidence that functionally impaired variants of the COQ2 gene (involved in the biosynthetic pathway for coenzyme Q10) are associated with an increased risk of developing MSA. This group previously identified multiplex families with MSA, indicating a genetic component in a disease that had previously been considered a non-genetic disorder.
Thursday, June 13, 2013
Best Post of November 2012: Photos reveal unique features of Einstein's cerebral cortex
The next in our "Best of the Month" series is from November 21, 2012:
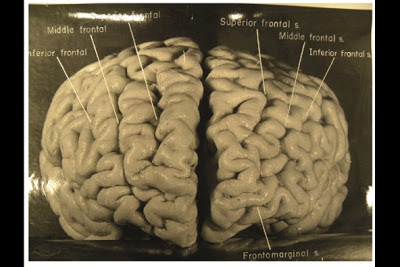
Photographs taken shortly after his death, but never before analyzed in detail, have now revealed that Einstein’s brain had several unusual features, providing clues about the neural basis of his extraordinary mental abilities.
Nature.com reports that, while doing Einstein's autopsy, the pathologist Thomas Harvey removed the physicist's brain and preserved it in formalin. He then took dozens of black and white photographs of it before it was cut up into 240 blocks. Now, anthropologist Dean Falk of Florida State University in Tallahassee and her colleagues have obtained 12 of Harvey’s original photographs from the National Museum of Health and Medicine in Silver Spring, Maryland, analyzed them, and compared the patterns of convoluted ridges and furrows with those of 85 brains described in other studies. Many of the photographs were taken from unusual angles, and show structures that were not visible in photographs that have been studied previously. The analysis was recently published in the journal Brain. The most striking observation, says Falk, was “the complexity and pattern of convolutions on certain parts of Einstein's cerebral cortex”, especially in the prefrontal cortex, and also parietal lobes and visual cortex.
Photographs taken shortly after his death, but never before analyzed in detail, have now revealed that Einstein’s brain had several unusual features, providing clues about the neural basis of his extraordinary mental abilities.
Nature.com reports that, while doing Einstein's autopsy, the pathologist Thomas Harvey removed the physicist's brain and preserved it in formalin. He then took dozens of black and white photographs of it before it was cut up into 240 blocks. Now, anthropologist Dean Falk of Florida State University in Tallahassee and her colleagues have obtained 12 of Harvey’s original photographs from the National Museum of Health and Medicine in Silver Spring, Maryland, analyzed them, and compared the patterns of convoluted ridges and furrows with those of 85 brains described in other studies. Many of the photographs were taken from unusual angles, and show structures that were not visible in photographs that have been studied previously. The analysis was recently published in the journal Brain. The most striking observation, says Falk, was “the complexity and pattern of convolutions on certain parts of Einstein's cerebral cortex”, especially in the prefrontal cortex, and also parietal lobes and visual cortex.
Friday, May 31, 2013
Best Post of October 2012: IDH1 Mutations in Oligodendrogliomas: PNA-clamping PCR is best analytic method
Time for the next in our "Best of the Month" series.. At the time of it's original publication, Dr. Craig Horbinski had this to say about a study which had recently been published in Brain Pathology: "Only catch is whether the method is TOO sensitive. After all, if all
oligos are positive for IDH1, why do the "inferior" IDH tests
effectively stratify prognosis? I'll be keen to see if anyone else can replicate this work."
Here's the original post from October 22, 2012:
Although direct sequencing of mutations in isocitrate dehydrogenase 1 (IDH1) has been considered to be the gold standard method to detect this mutation, the sensitivity of this technique has been questioned especially because specimens from glial tumors may contain large numbers of non-tumor cells. The group screened 141 cases of oligodendroglial tumors for IDH1 mutations using peptide nucleic acid (PNA)-mediated clamping PCR and compared the results with the results of direct sequencing, pyrosequencing, and immunohistochemistry (IHC). Nested PCR was only performed in cases having mutant IDH1 only discovered by clamping PCR. Using dilution experiments mixing IDH1 wild-type and mutant DNA samples, clamping PCR detected mutations in samples with a 1% tumor DNA composition. Using PNA clamping PCR, the group detected 138 of 141 (97.9%) cases with mutant IDH1 in the series, which is significantly higher (P = 0.016; PNA clamping vs. direct sequencing) than those of direct sequencing (74.5%), pyrosequencing (75.2%), and IHC (75.9%). From these results, it appears that almost all oligodendroglial tumors have IDH1 mutations, suggesting that IDH1 mutation is an early and common event especially in the development of oligodendroglial tumors.
Here's the original post from October 22, 2012:
Although direct sequencing of mutations in isocitrate dehydrogenase 1 (IDH1) has been considered to be the gold standard method to detect this mutation, the sensitivity of this technique has been questioned especially because specimens from glial tumors may contain large numbers of non-tumor cells. The group screened 141 cases of oligodendroglial tumors for IDH1 mutations using peptide nucleic acid (PNA)-mediated clamping PCR and compared the results with the results of direct sequencing, pyrosequencing, and immunohistochemistry (IHC). Nested PCR was only performed in cases having mutant IDH1 only discovered by clamping PCR. Using dilution experiments mixing IDH1 wild-type and mutant DNA samples, clamping PCR detected mutations in samples with a 1% tumor DNA composition. Using PNA clamping PCR, the group detected 138 of 141 (97.9%) cases with mutant IDH1 in the series, which is significantly higher (P = 0.016; PNA clamping vs. direct sequencing) than those of direct sequencing (74.5%), pyrosequencing (75.2%), and IHC (75.9%). From these results, it appears that almost all oligodendroglial tumors have IDH1 mutations, suggesting that IDH1 mutation is an early and common event especially in the development of oligodendroglial tumors.
Saturday, May 18, 2013
Neuropathologist Roger Brumback found murdered in his home
Neuropathologist Roger Brumback, 65, and his wife were found murdered in their Nebraska home on Tuesday. Dr. Brumback, an active member of the neuropathology community, was the former chair of pathology at Creighton University in Omaha.
| Roger Brumback, MD |
The bodies of Dr. Brumback and his wife, Mary, were discovered by a piano mover who had arrived at their home to find the front door ajar.
In addition to his work as a neuropathologist, Dr. Brumback was a clinical pediatric neurologist and was the founding Editor-in-Chief of the Journal of Child Neurology. At a memorial held at Creighton in his honor, medical students described Brumback as a kind and caring man who was devoted to teaching.
Police are looking at any link between this killing and that of the son of another Creighton pathologist back in 2008. There have been no arrests in either case.
Tuesday, May 14, 2013
Dr. Mark Cohen's "Wicked Clown" Mitochondrion
Inspired by the last post regarding unusual mitochondrial inclusions, the insurmountable Dr. Mark Cohen of Case Western sent in his favorite picture of an aberrant mitochondrion, which he likens to a "wicked clown":
Monday, May 6, 2013
Angulated intramitochondrial inclusions in the left leg of a man with a left-sided limp
A 59-year-old male with a one-year history of limping as well as pain, weakness, and paresthesia of the left lower extremity who underwent lumbar microdiscectomy with cord decompression. Although the patient's pain subsided post-operatively, his other symptoms persisted. Since his CPK levels were chronically elevated (around the 500's), biopsies were later performed on the quadriceps bilaterally. Light microscopic examination was underwhelming except for some denervation effect on the left. However, subsarcolemmal clumping was evident with NADH histochemisty on both the right and left. This clumping prompted me to perform an ultrastructural examination, which revealed the following angulated intramitochondrial inclusions IN THE WEAK LEG ONLY:
These inclusions were present in about 5% of mitochondria. I reported out my findings in a descriptive manner, not certain of their significance. If anyone has seen such intramitochondrial inclusions in a specimen, please comment. As of now, the patient has no definitive diagnosis.
These inclusions were present in about 5% of mitochondria. I reported out my findings in a descriptive manner, not certain of their significance. If anyone has seen such intramitochondrial inclusions in a specimen, please comment. As of now, the patient has no definitive diagnosis.
Friday, April 26, 2013
Annual AANP meeting is two months away
 |
| The Charleston Place Hotel in Charleston, South Carolina |
Tuesday, April 16, 2013
A Rich Focus
 |
| The grey arrow is pointing to a Rich focus that has hemorrhaged into the subarachoid space |
Tuesday, April 9, 2013
Best Post of September 2012: DecisionDx-GBM: Should every glioblastoma patient be getting this test?
The next in our Best of the Month Series is from September 12, 2012. It's worth revisiting the question of whether or not genetic testing of brain tumors is appropriate in patients who are not on research studies:
What does the neuropathology community think of DecisionDx-GBM? This is a product offered by Castle Biosciences, based in Phoenix, AZ. DecisionDx-GBM is a gene expression profile test developed at The University of Texas M. D. Anderson Cancer Center for the purpose of increasing the accuracy of the prognosis and predicted responsiveness of glioblastoma multiforme to first line radiation plus temozolomide. The test is able to distinguish GBM tumors with a proneural phenotype (tumor signature) from those with a mesenchymal / angiogenic phenotype. Patients with a proneural phenotype tumor who are treated with first line radiation plus temozolomide experience a significantly longer median survival (over 7 years) compared to those patients with a mesenchymal / angiogenic phenotype tumor (approximately 1 year). According to the company website, the assay has been fully validated and has been available for clinical use since 2008. A study is ongoing to determine whether the tumor molecular profile conferring a mesenchymal/angiogenic phenotype is associated with a selective increase in benefit from the addition of bevacizumab to temozolomide and radiotherapy. DecisionDx-GBM is also currently being incorporated into a number of other prospective and retrospective studies.
Is DecisionDx-GBM covered by insurance? Castle Biosciences states that the test receives reimbursement from a number of commercial insurance companies, and appeals to the Administrative Law Judge level for Medicare have resulted in favorable decisions for full payment. Should a patient need to pay out-of-pocket for this test, I could not find on the website what the cost would be.
Should we as neuropathologists recommend the use of this profile? Here is a an example of the kind of report that is generated when one orders the panel of 12 genes (3 of which are for control) upon sending a block of paraffin-embedded fixed tissue to the company. I would be interested in hearing people's opinions regarding this product, and whether there are others on the market which might be comparable. Please post!
Thursday, March 21, 2013
How do you calculate a brain tumor's MIB-1 index?
 |
| Elizabeth J. Cochran, MD |
I was recently contacted by the affable Elizabeth Cochran, MD asking if I would post a query regarding MIB-1 counts on brain tumors. Does anyone know if there is a standardized approach to this? Please tell us your approach to MIB-1 quantification in the comment section. Dr. Cochran takes an approach based on an article she had read in Human Pathology some time ago, summarized as follows:
1. Examine control section stained with MIB-1 antibody to
confirm that the stain worked.
2. Survey the slide of the case to be counted to find the
area(s) that have the most stained nuclei.
3. Put reticule in right eyepiece of microscope.
4. Place the area identified in #2 under the reticule at 40x
power.
5. Using the hand held counter, count all the immunoreactive
nuclei within the entire grid.
6. Count all nuclei that touch the right and bottom outer
border of the grid, but NOT the nuclei that touch the upper and left borders.
7. Record this number in the lab book.
8. Count all nuclei present within one vertical column of
the grid. This column should be randomly
chosen.
9. Record this number in the lab book.
10. Count all nuclei present within one horizontal column of
the grid. Again this column should be
randomly chosen, and recorded in the lab book.
11. Again, be sure to count all nuclei that touch the right
and bottom outer border of the grid, but NOT the nuclei that touch the upper
and left borders
12. Repeat this procedure for at least two more high power
fields on the section that contains the most numerous immunostained nuclei and
record all values in the lab book.
13. Calculate the
MIB-1 proliferative index as follows:
a. For
each of the fields counted: Add
all of the nuclei counted in each row and column from each field and multiple
by five. Divide the number of the
immunostained nuclei counted in that field by the calculated sum of all of the
nuclei in that field.
b. The
total number of all cells counted in all fields should be >
1000. If it is substantially less than
1000, count another field, as described above.
c.
Calculate the average of the three fields and
record the proliferative index in the lab book.
Subscribe to:
Comments (Atom)
Neuropathology Blog is Signing Off
Neuropathology Blog has run its course. It's been a fantastic experience authoring this blog over many years. The blog has been a source...